Linear Regression
The Linear Regression tab performs ordinary least squares (OLS) linear regression analysis. OLS finds the coefficient vector that minimizes the residual sum of squares. See Regression Analysis Fundamentals for the mathematical background.
For count data, binary outcomes, or other non-normal response variables, use Generalized Linear Model (GLM) instead.
Basic Usage
Opening Linear Regression
Select Analysis > Linear Regression (OLS)... from the menu bar to open a new Linear Regression tab.
Setting Up Variables
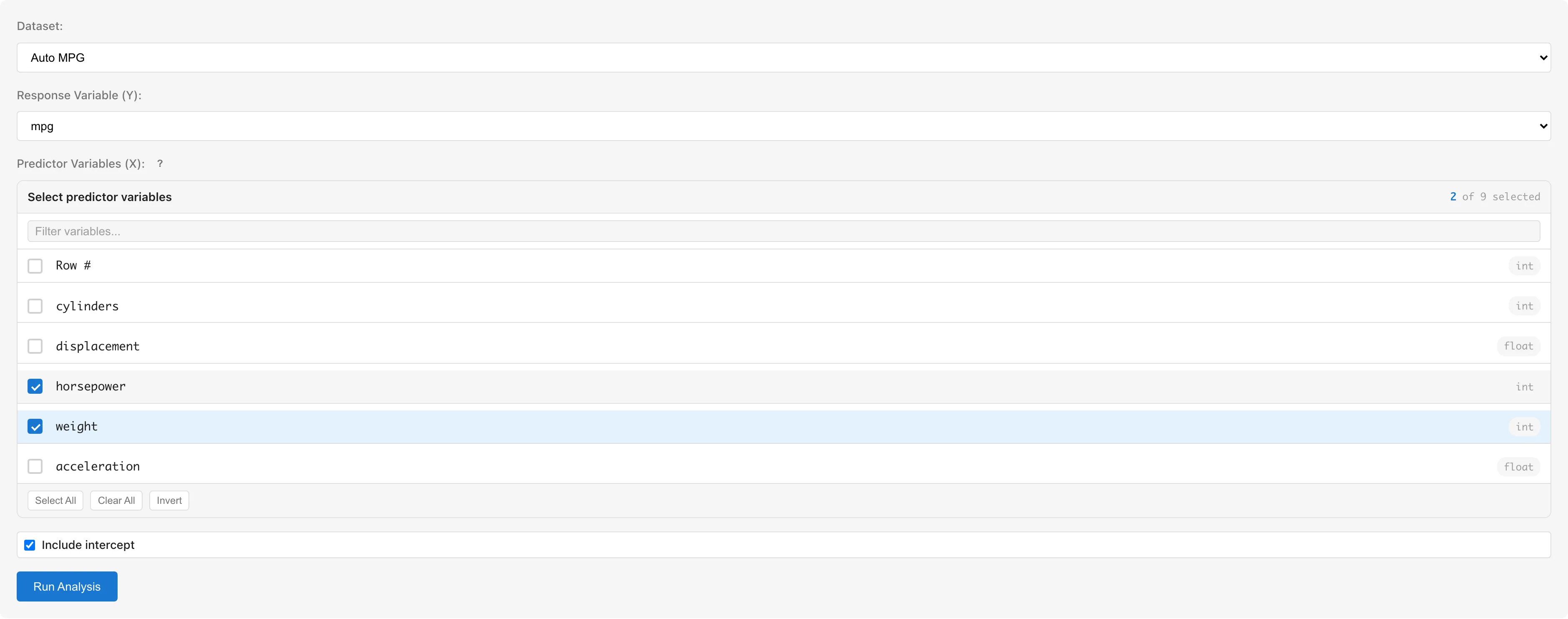
Dataset selects the dataset to analyze.
Response Variable (Y) selects the response variable. Only numeric columns (int64, float64) are available.
Predictor Variables (X) selects predictor variables using checkboxes. Only numeric columns are selectable; non-numeric columns are grayed out. To use categorical variables, convert them to numeric dummy variables using the Dummy Coding tab first (see Notes).
Include intercept toggles the intercept term. Enabled by default.
Click the Run Analysis button to run the analysis.
Understanding Results
Model Summary
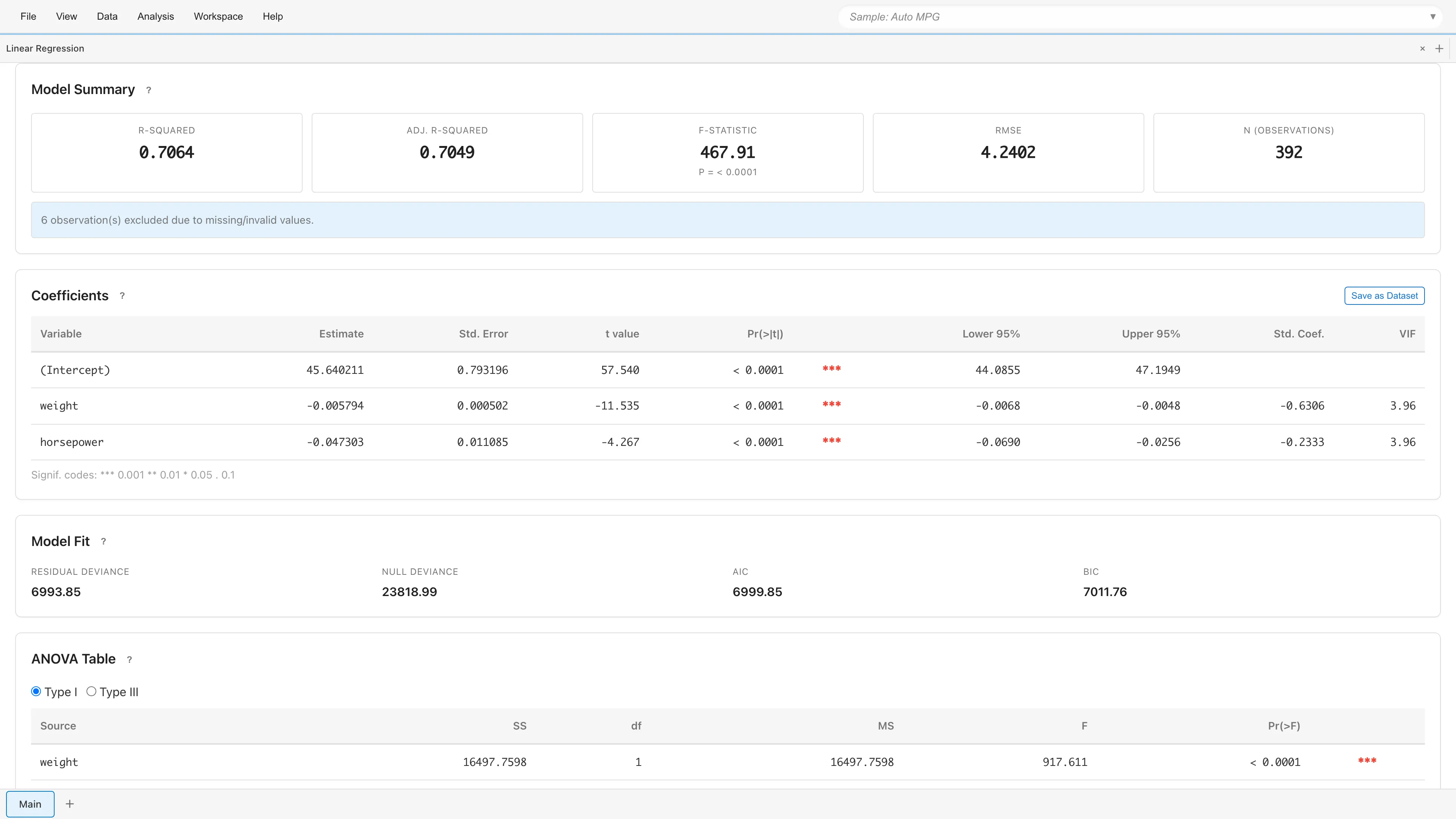
Displays overall model fit statistics.
| Metric | Description |
|---|---|
| R-squared | Proportion of variance explained by the model (0 to 1) |
| Adj. R-squared | R-squared adjusted for the number of predictors |
| F-statistic | Overall model significance test (with p-value) |
| RMSE | Standard deviation of residuals (measure of prediction error) |
| N (observations) | Number of observations used in the analysis |
If rows containing missing or invalid values were excluded, the number of excluded rows is displayed.
Coefficients
Displays regression coefficients and test results for each variable.
| Column | Description |
|---|---|
| Variable | Variable name (intercept shown as "(Intercept)") |
| Estimate | Estimated coefficient |
| Std. Error | Standard error |
| t value | -statistic , following a distribution |
| Pr(>|t|) | Two-sided p-value from the distribution |
| (Significance marks) | *** p<0.001, ** p<0.01, * p<0.05, . p<0.1 |
| Lower 95% / Upper 95% | 95% confidence interval |
| Std. Coef. | Standardized coefficient for comparing effect sizes across variables (N/A for intercept) |
| VIF | Variance Inflation Factor (see Multicollinearity) |
Under normal errors, the OLS -test is exact regardless of sample size.
The coefficients table can be saved as a dataset using the Save as Dataset button for export to CSV.
Interpreting Coefficients
Coefficients are directly interpretable on the response scale.
- Continuous predictor: Holding other variables constant, a one-unit increase in changes by
- Dummy variable: Represents the difference in relative to the reference category
- Intercept: when all predictors are zero
- Standardized coefficient (Std. Coef.): Expressed in standard deviation units, enabling direct comparison of effect sizes across variables with different scales
Model Fit
| Metric | Description |
|---|---|
| Residual Deviance | Residual sum of squares |
| Null Deviance | Total sum of squares |
| AIC | Akaike Information Criterion . Lower is better |
| BIC | Bayesian Information Criterion . Penalizes complexity more strongly than AIC |
ANOVA Table
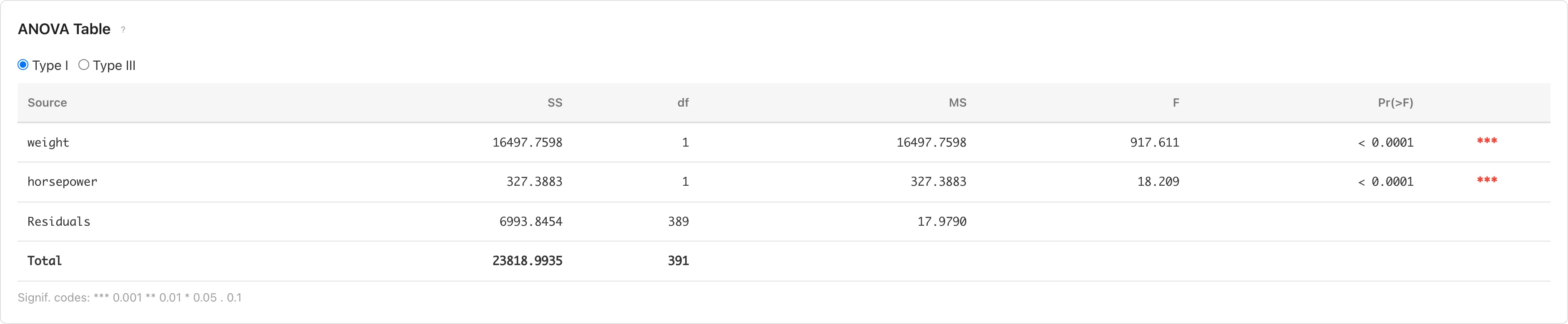
Evaluates each predictor's contribution through analysis of variance. Switch between Type I and Type III using radio buttons.
- Type I (Sequential): Computes sum of squares in the order variables were entered. Results depend on variable entry order.
- Type III (Partial): Computes sum of squares as if each variable were entered last. Results are independent of variable entry order.
| Column | Description |
|---|---|
| Source | Source of variation |
| SS | Sum of squares |
| df | Degrees of freedom |
| MS | Mean square (SS / df) |
| F | F-statistic (MS / MS_Residual) |
| Pr(>F) | p-value based on the F distribution |
Prediction & Confidence Intervals
Displays predicted values and interval estimates for each observation.
| Column | Description |
|---|---|
| Obs | Observation number |
| Fitted | Predicted value |
| 95% CI Lower / Upper | 95% confidence interval for the mean response |
| 95% PI Lower / Upper | 95% prediction interval for individual observations |
The confidence interval (CI) represents the precision of the estimated mean, while the prediction interval (PI) represents the range for a new individual observation. PI is always wider than CI. The confidence level is fixed at 95%.
When exceeding 100 rows, only the first 50 rows are displayed. Click Show all N rows to display all rows.
Saving and Diagnostics

Save analysis results to the project and view diagnostic plots.
Saving the Model
Enter a model name in the Model Name field and click Save Model. The model name defaults to the format "Linear Regression: Y ~ X1 + X2".
If an existing model with the same configuration (dataset, response variable, predictor variables) exists, a confirmation dialog for overwriting is displayed.
Data Generated on Save
Saving a model automatically creates a derived dataset that adds diagnostic columns to the original data.
| Column | Symbol | Description |
|---|---|---|
fitted_values | Predicted values | |
deviance_residuals | Residuals | |
standardized_residuals | Standardized residuals | |
leverage | Leverage (diagonal of the hat matrix) | |
cooks_distance | Cook's Distance |
Diagnostics and Details
After saving the model, two buttons appear:
- View Model Details - Opens the Model Detail tab showing detailed model information
- View Diagnostics - Opens the Residual Diagnostics tab showing residual diagnostic plots
Residual Diagnostics
View Diagnostics opens four diagnostic plots for assessing the OLS assumptions:
OLS assumptions:
- Linearity - A linear relationship exists between the response and predictor variables
- Normality - Residuals follow a normal distribution
- Homoscedasticity - The variance of residuals is constant and does not depend on fitted values
- Independence - Residuals are independent of each other (not directly verifiable through diagnostic plots). For hierarchical or clustered data, consider using GLMM with random effects. MIDAS does not currently support time series autocorrelation
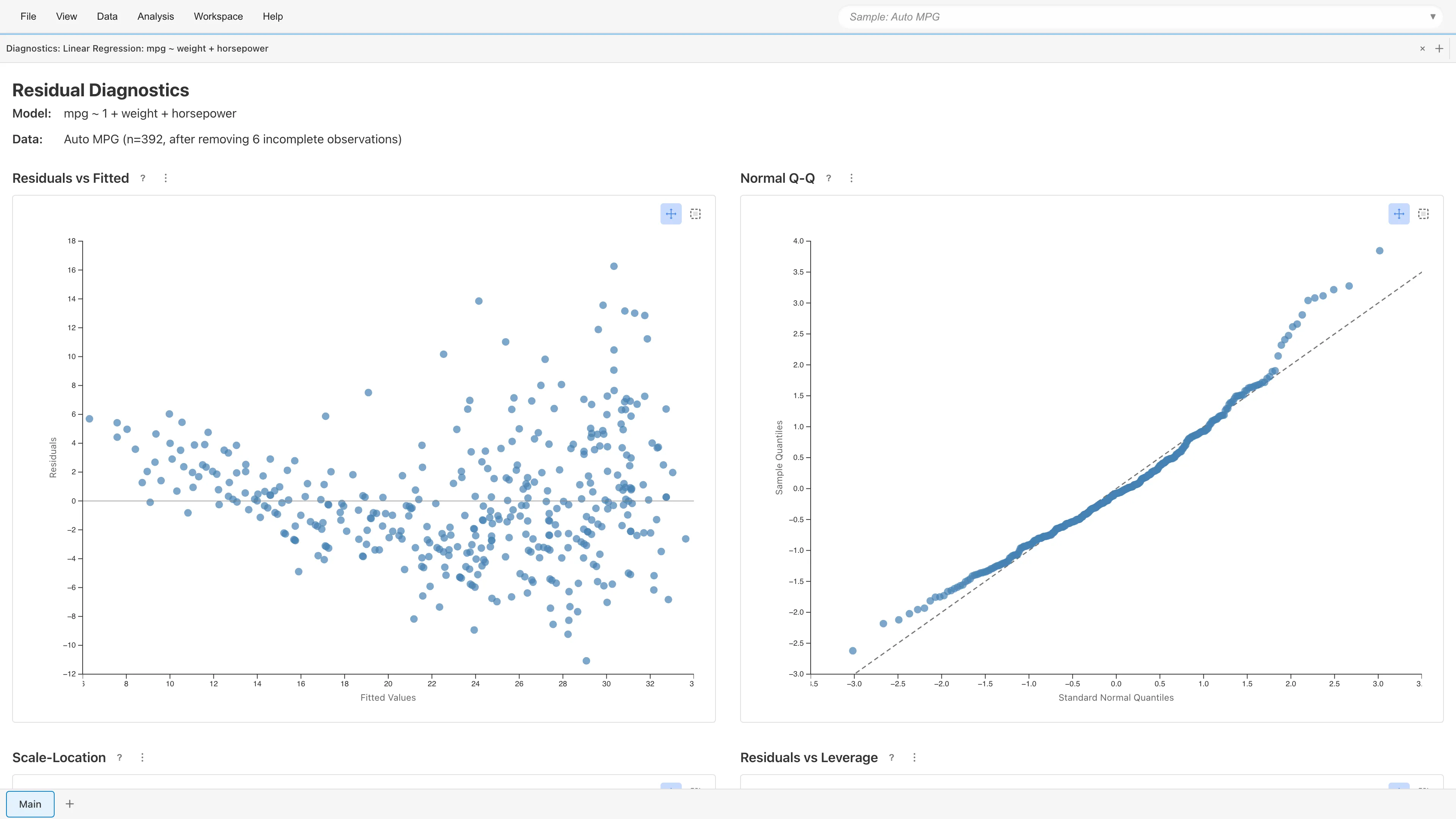
The diagnostic plots use standardized residuals . See Regression Analysis Fundamentals for the formulas.
Residuals vs Fitted
Plots residuals against fitted values . With a well-specified model, residuals scatter randomly around zero.
- Curved pattern: Nonlinear effects of predictors may be missing
- Funnel-shaped pattern: Possible heteroscedasticity (check Scale-Location plot for details)
Normal Q-Q Plot
Plots standardized residual quantiles against theoretical normal quantiles. Points fall along the diagonal when residuals are normally distributed. Deviation at both ends indicates heavy tails; an S-shaped departure indicates skewness.
Scale-Location
Plots against fitted values. Constant variance produces an even horizontal spread. A funnel-shaped or upward-trending pattern indicates heteroscedasticity. When heteroscedasticity is present, coefficient estimates remain unbiased but standard errors become inaccurate, making confidence intervals and p-values unreliable.
Residuals vs Leverage
Plots standardized residuals against leverage . Cook's (1977) distance contours (: orange dashed, : red dashed) are overlaid. Cook suggested comparing to the 50th percentile of , which is typically near 1. This is not a formal rejection threshold but a guide for comparing relative influence among observations.
- Leverage: Measures how far an observation's predictor values are from others. is the conventional threshold for high leverage
- Cook's Distance: Combines leverage and residual size into a single influence measure
Observations outside the contour lines may substantially change the model estimates if removed.
Point Selection
Click or rectangle-select data points on any plot to display details (fitted values, residuals, leverage, Cook's Distance, etc.) in a table below the plots. Selection state is synchronized across all four plots.
Notes
Using Categorical Variables
OLS regression only accepts numeric variables. To use string or boolean categorical variables as predictors, convert them to numeric dummy variables using the Dummy Coding tab before running the analysis.
Automatic Exclusion of Missing and Invalid Values
Rows containing missing values (null), non-numeric values, or infinity are automatically excluded from the analysis. The number of excluded rows is displayed in the Model Summary.
Multicollinearity
When predictors are highly correlated, coefficient estimates become unstable. If VIF (Variance Inflation Factor) is large for any variable in the coefficients table, consider removing redundant variables or combining correlated predictors. See Regression Analysis Fundamentals for details on VIF.
Sample Size and Normality
The finite-sample exactness of -tests and -tests depends on the normality of errors. With large samples, the central limit theorem ensures the test statistics are approximately normal, but for small samples, verify residual normality using the Q-Q plot. The required sample size depends on the true error distribution, so no universal threshold applies.
References
- Cook, R. D. (1977). Detection of influential observation in linear regression. Technometrics, 19(1), 15-18.
Next steps
- Residual Diagnostics - Verify model assumptions
- Generalized Linear Model (GLM) - Modeling count data and binary outcomes
See also
- Regression Analysis Fundamentals - Mathematical background of OLS and diagnostic statistics
- Dummy Coding - Converting categorical variables to numeric