ANOVA (Analysis of Variance)
The ANOVA tab analyzes whether the means of a response variable differ across groups defined by categorical variables. Both one-way and two-way designs are supported.
Basic Usage
Open the Tab
Select Analysis > ANOVA... from the menu bar.
Run an Analysis
Configure the following in the settings panel:
- Select a dataset from Dataset
- Choose One-Way or Two-Way under Analysis Type
- Select a categorical variable for Factor A
- Select a numeric variable for Response Variable
- Click Run Analysis
Data Format
Data must be in long format with one row per observation. Each row contains the factor value and the response variable value. Use Reshape to convert wide-format data.
One-Way ANOVA
Analyzes differences in the response variable means across groups defined by a single categorical factor. Use this when you have one grouping factor.
Statistical Model
is the -th observation in group , is the overall mean, is the effect of group , and is the error term.
Null Hypothesis
Tests whether all group population means are equal.
Variable Selection
Factor A: Select a categorical variable that defines the groups. Columns with nominal or ordinal measurement scale appear as options.
Response Variable: Select the numeric variable to analyze. Columns with interval or ratio measurement scale appear as options.
Example
To analyze whether sepal length differs among the three Iris species (setosa, versicolor, virginica) in the Iris sample data:
- Dataset: Iris
- Analysis Type: One-Way
- Factor A:
species - Response Variable:
sepal_length - Click Run Analysis
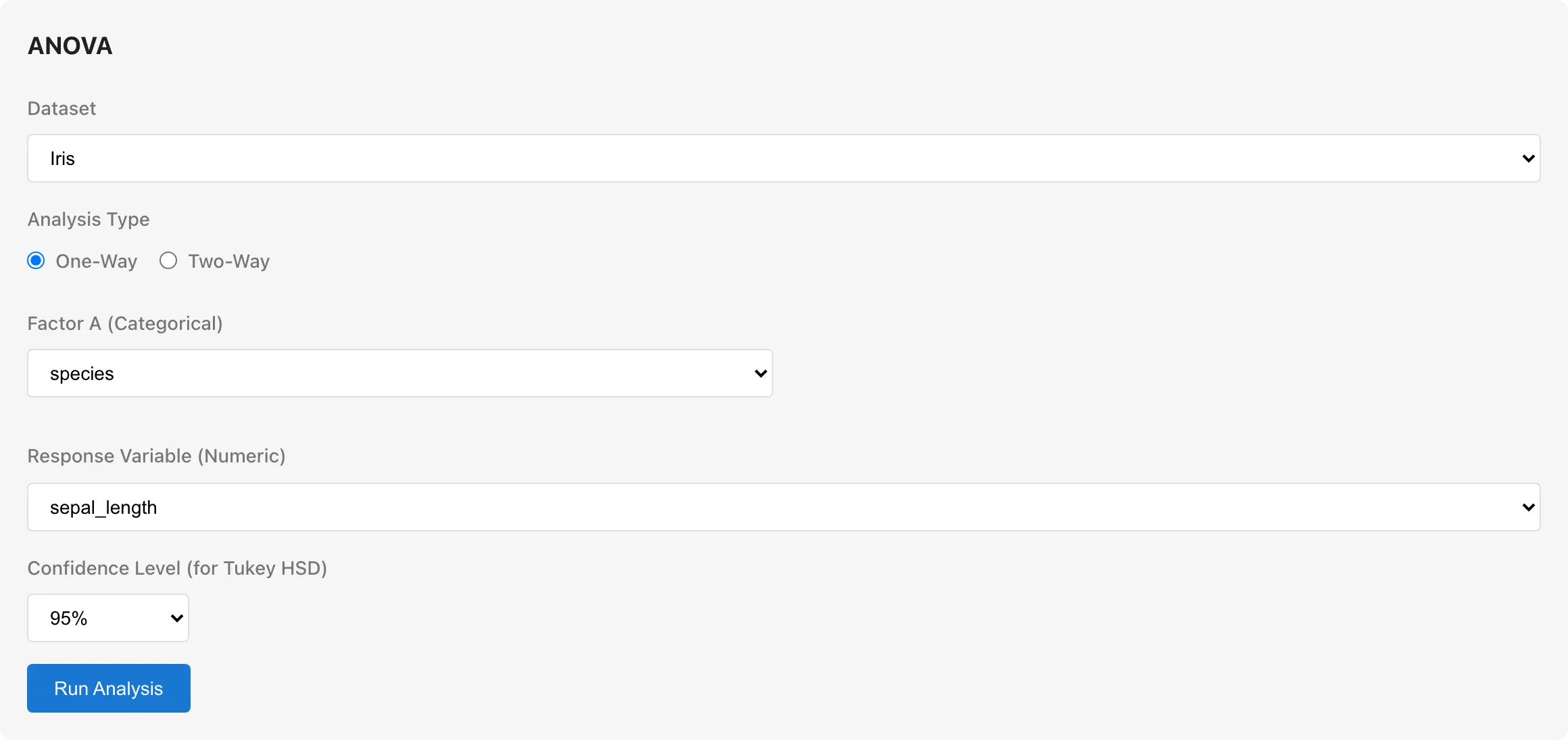
Confidence Level
Set the confidence level for the F-test and Tukey HSD post-hoc comparisons. Choose from 90%, 95% (default), or 99%. The significance level α equals 1 − confidence level (95% corresponds to α = 0.05). The Tukey HSD confidence interval width matches the confidence level.
Two-Way ANOVA
Analyzes the effects of two categorical factors and their interaction on the response variable. Use this when you have two grouping factors.
Statistical Model
With interaction:
is the effect of factor A, is the effect of factor B, and is the interaction effect.
Additional Settings
Factor B: Select a second categorical variable, different from Factor A.
Include interaction term (A x B): Whether to include the interaction term in the model. Enabled by default. Include the interaction when the effect of one factor may depend on the level of the other. If the interaction is known to be absent, excluding it increases the residual degrees of freedom, providing a more precise estimate of the error variance.
Sum of Squares Type: Choose the method for computing sums of squares.
Sum of Squares Types
Type I computes sums of squares sequentially based on the order factors enter the model. Each factor's contribution depends on which factors are already in the model. MIDAS enters Factor A first, then Factor B, then the interaction term. The SS for Factor A is computed without considering other factors, and the SS for Factor B reflects its contribution after removing the effect of Factor A. Swapping the Factor A and Factor B assignments changes the results.
Type III computes sums of squares for each factor as if it were the last one entered. Each factor's contribution is adjusted for all other factors.
For balanced designs (equal sample sizes in all cells), Type I and Type III produce identical results. For unbalanced designs, Type III is generally preferred because results do not depend on factor ordering.
Type III Interpretation with Interaction
MIDAS uses treatment coding. Treatment coding designates one level of each factor as the reference category (baseline level) and expresses the effects of all other levels as differences from it. The reference category is the first level in alphabetical order. When the interaction term is included, the Type III test for a main effect estimates the effect of that factor while the other factor is at its reference level. For example, if Factor A has levels A, B, C and Factor B has levels X, Y, the reference categories are A and X, and the Type III test for the main effect of Factor A tests the effect of Factor A when Factor B is at X. With balanced data, this coincides with the test about marginal means averaged across all levels. With unbalanced data, the two may differ.
Reading the Results
Observations
The total number of observations used in the analysis appears at the top. If rows were excluded due to missing values, the count of excluded rows is also shown.
Group Statistics
A summary table of descriptive statistics for each group.
| Column | Description |
|---|---|
| Group | Group name |
| N | Number of observations |
| Mean | Group mean |
| SD | Standard deviation (square root of unbiased variance, denominator n − 1) |
| CI Lower / CI Upper | Confidence interval for the group mean. The confidence level matches the Confidence Level setting |
| Min | Minimum value |
| Max | Maximum value |
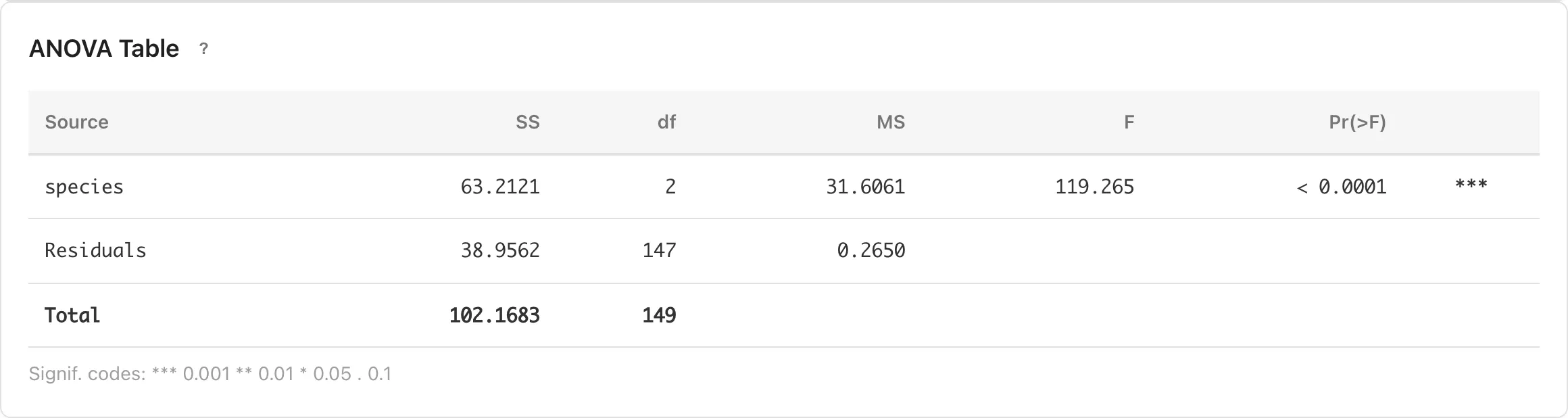
ANOVA Table
The main results table. Decomposes the total variance of the response variable into contributions from each factor and residual error.
| Column | Description |
|---|---|
| Source | Source of variation |
| SS | Sum of squares -- the amount of variation attributable to each source |
| df | Degrees of freedom |
| MS | Mean square (SS / df) |
| F | F statistic (MS of the source / MS of residuals) |
| Pr(>F) | p-value -- the probability of observing an F statistic as extreme as, or more extreme than, the observed value under the null hypothesis |
| Partial η² | Partial eta-squared. Computed as SS_effect / (SS_effect + SS_residual). Measures the proportion of variance accounted for by the source relative to the source and residual variance combined |
| Partial ω² | Partial omega-squared. A degrees-of-freedom-adjusted effect size estimator with less upward bias than partial η² for estimating the population effect size. Displayed as 0 when the estimate is negative |
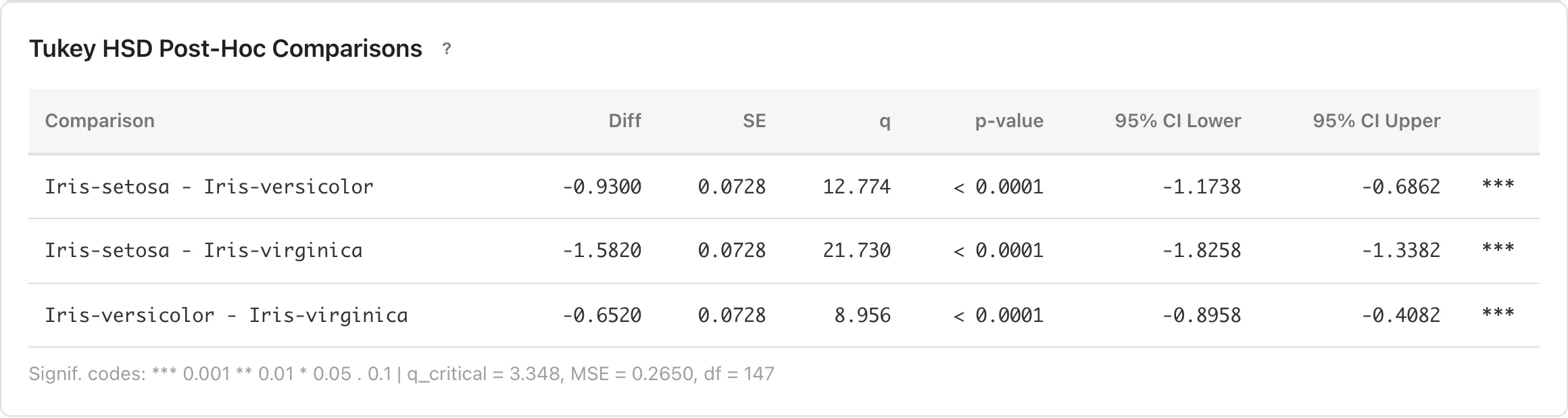
Tukey HSD Post-Hoc Comparisons
The ANOVA F-test determines whether at least one group mean differs from the others, but does not show how large the difference is for each pair. Tukey HSD post-hoc comparisons estimate the mean difference and its simultaneous confidence interval for every pair of groups, allowing you to assess the magnitude and precision of each pairwise difference.
For one-way ANOVA, Tukey HSD is computed automatically regardless of the F-test result. The Tukey-Kramer method is used, which handles unequal group sizes.
Tukey HSD constructs simultaneous confidence intervals for all pairwise mean differences. It controls the family-wise error rate, reducing the inflation of false positives compared to running individual t-tests for each pair.
| Column | Description |
|---|---|
| Comparison | The two groups being compared |
| Diff | Difference in means (Group 1 mean − Group 2 mean) |
| SE | Standard error of the difference |
| q | Studentized range statistic |
| p-value | p-value from the studentized range distribution |
| CI Lower / CI Upper | Simultaneous confidence interval for the mean difference. Adjusted so that all pairwise intervals simultaneously contain the true values with at least the specified confidence level |
The critical value , MSE, and residual degrees of freedom are displayed below the table.
Assumptions
ANOVA assumes the following. Verify that these are reasonable when interpreting results.
- Independence: Observations are independent of each other
- Normality: The response variable follows a normal distribution within each group. With large sample sizes, the central limit theorem causes the sampling distribution of group means to approach normality, so the Type I error rate of the F-test is less likely to deviate substantially from the nominal level
- Homogeneity of variance: The variance is equal across all groups
Assumption Diagnostics
A residual Q-Q plot is displayed below the ANOVA table. The Q-Q plot compares the distribution of residuals against a theoretical normal distribution. Points falling close to the diagonal line suggest approximate normality. Departures such as heavy tails or skewness are visible in the shape of the deviation from the line.
Homogeneity of variance can be assessed by comparing the SD values in the Group Statistics table. If the standard deviations differ substantially across groups, interpret the ANOVA results with caution.
In two-way ANOVA, residuals are computed from the fitted values of the selected model. When the interaction term is included, the model fits a separate mean for each cell, so residuals equal deviations from cell means. When the interaction term is excluded, the main-effects-only model produces different fitted values, and the residuals reflect deviations from those predictions. The choice of model therefore affects the Q-Q plot.
Error Messages
In two-way ANOVA, if any combination of factor levels has no observations, the model with interaction cannot be estimated. The error "The design matrix is rank deficient" is displayed. Turn off the interaction term or check whether your data has empty cells.
Missing Values
Rows containing missing values are automatically excluded. The number of excluded rows is displayed in the results panel. For two-way ANOVA, rows with missing values in either factor or the response variable are excluded.
Related Pages
- Linear Regression -- the ANOVA table in the regression tab tests the overall model fit, while this tab uses categorical factors